Tepidimonas/Cloacibacterium
Specific identification and quantification of the bacteria indicator
groups Tepidimonas und Cloacibacterium in paper mill cycles
Item Number: 01120020
Lab analysis: detection of biofilm forming bacteria
In paper mills, monitoring and directed control and avoidance of biofilms play a big role. Basically, indicator organisms for early biofilm formation are bacteria of the genus Tepidimonas and Cloacibacterium. We analyze your sample for this bacteria specifically, reliably and fast – and quantify the microorganisms for a real risk assessment or for cleansing controls. Our analysis is your early warning system for biofilm formation.
Qualitative or quantitative detection?
Quantitative detection
Which method is applied?
The reliable VIT® gene probe technology with specific gene probes. Results will be provided already within 3 working days after sample receipt.
Which requirements are important?
- Water samples: quantitative analysis, minimum 100 ml
- Biofilm: Please request our biofilm sampling protocol
Alternatively, for analysis directly on site, we also offer the test kits
VIT® Cloacibacterium
VIT® Tepidimonas
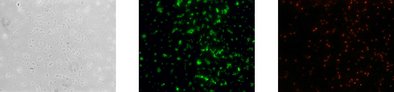
Your Advantages
reliable detection and quantification per filtrated volume of bacteria of the genera Tepidimonas and Cloacibacterium
cultivation-independent analysis
only living bacteria are detected
maximum specificity
early recognition of biofilm formation due to indicator function of detected microorganisms