Bio-P Analysis
Quantitative analysis of polyphosphate-accumulating organisms (PAO)
and glycogen-accumulating organisms (GAO)
Item Number: 01120014
Lab analysis: monitoring the biological phosphate removal
Identification and quantification of the bacteria responsible for EBPR processes. Specific gene probes detect the polyphosphate-accumulating organisms (PAO), as well as their counterpart, the glycogen-accumulating organisms (GAO). For both populations, the quantitative results are provided as % shares of the living bacteria flora.
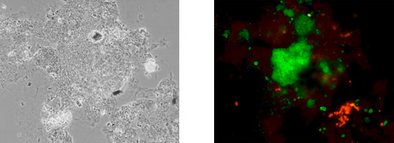
Representative photo documentation of the different EBPR organisms visualized directly in the sample material is added to the report.
Qualitative or quantitative detection?
Quantitative detection
Which method is applied?
The reliable VIT® gene probe technology.
Which requirements are important?
Please request our simple protocol for fixation of wastewater samples. Make sure that you fixate/stabilize your samples in the indicated manner before sending the samples to us.
For an analysis directly on site, we recommend our test kit VIT® PAO/GAO.
Your Advantages
specific detection of Bio-P bacteria (PAO and GAO)
quantification of populations directly in samples
detection of living bacteria exclusively