Nostocoida limicola II
Identification and quantification of the filamentous bacteria
Nostocoida limicola Typ II and Alysiosphaera in wastewater samples
Item Number: 01130007
Lab analysis: detection of filamentous bacteria
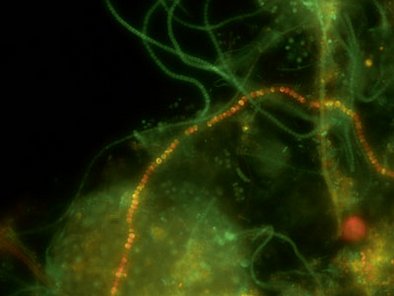
Identification and quantification of Gram-positive Nostocoida limicola type II as well as Gram-negative Alysiosphaera filaments with our VIT® Nostocoida limicola II test kit. The two pearl necklace-formed filaments are morphologically very similar so conventionally the distinction is not possible. Though specific gene probes typify and quantifiy them unequivocally.
The results are displayed according to the VIT® key. The report contains a significant photo documentation that visualises the filaments in the wastewater sample.
Qualitative or quantitative detection?
Semi-quantitative detection
Which analysis is applied?
The VIT® gene probe technology.
Which requirements are important?
Please request our simple protocol for fixation of wastewater samples. Make sure that you fixate/stabilize your samples in the indicated manner before sending the samples to us.
For an on-site analysis, we recommend our test kit VIT® Nostocoida limicola II.
Your Advantages
specific detection of Nostocoida limicola type II and Alysiosphaera
detection of living bacteria exclusively
quantification of filaments even in dense sludge flocs
highest specificity
applied for the prevention of disturbances