Acetic acid bacteria
Qualitative detection of the bacteria genera Acetobacter, Asaia,
Gluconoacetobacter and Gluconobacter in alcoholic beverages and soft drinks
Item Number: 01220003
Lab analysis: detection of beverage spoilage acetic acid bacteria
Alcoholic as well as non-alcoholic beverages will be analyzed for presence of following genera of beverage spoiling acetic acid bacteria: Acetobacter, Asaia, Gluconoacetobacter and Gluconobacter.
Results will be provided within three working days after sample receipt, in case of readily enriched samples or grown colonies by the next working day.
Qualitative or quantitative detection?
Qualitative
Which method is applied?
The reliable VIT® gene probe technology.
Which requirements are important?
Product or raw material samples:
- beer, wine, juices and soft drinks, minimum 100 ml
- juice concentrates, minimum 25 ml
- water samples (e.g. soda, rinsing water etc.), minimum 150 ml
Pre-enriched samples:
- Pre-enriched bouillons (10 ml) or agar plates with colonies: This allows an acceleration of time-to-result (results are provided within 1 working day after sample receipt).
Detection of acetic acid bacteria is also part of our service Screening "Beverage-spoiling microorganisms".
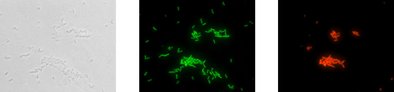
Your Advantages
specific detection of acetic acid bacteria relevant for beverage industry
detection of living bacteria exclusively
short operational time
highest specificity
analysis of diverse sample material